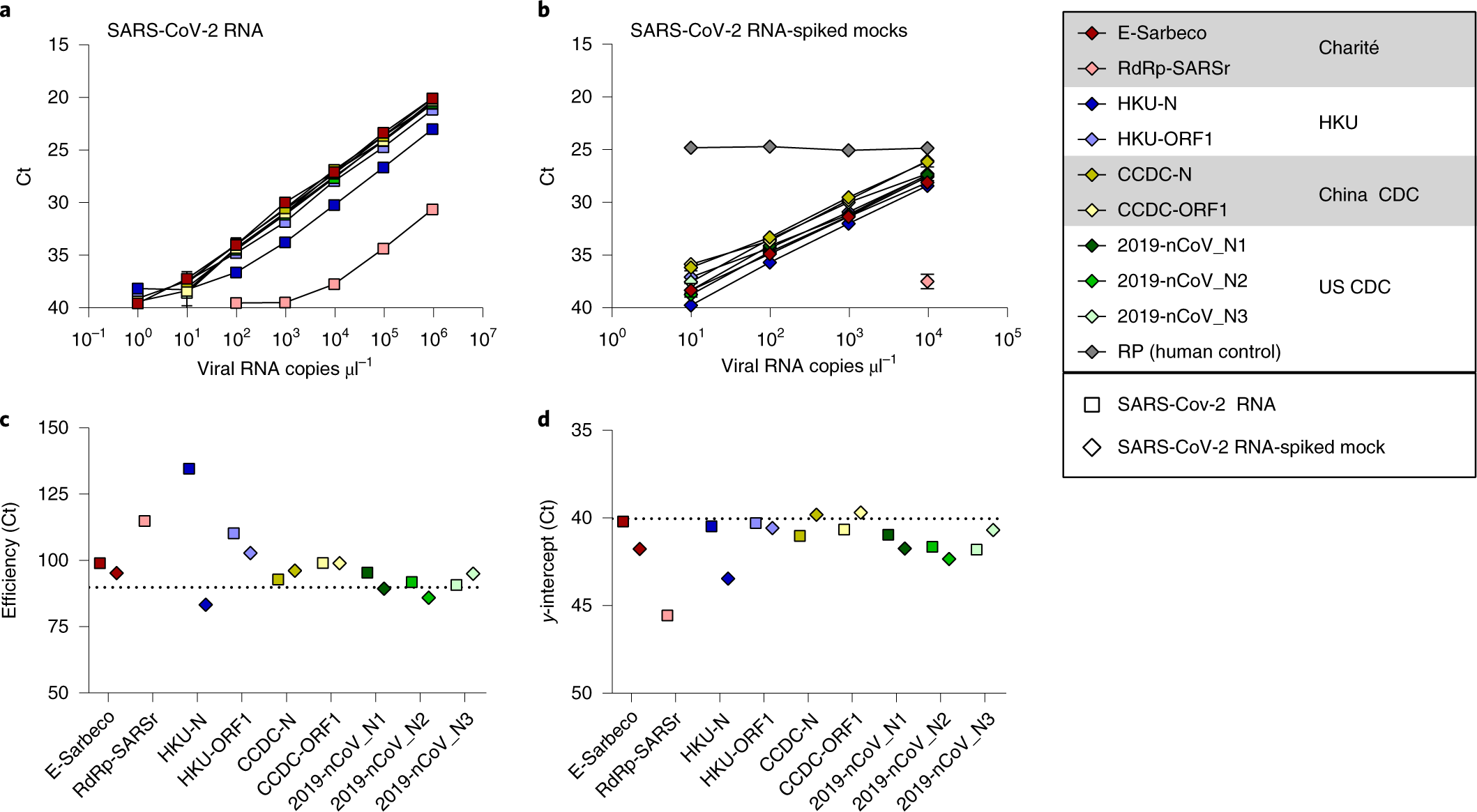
Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT–qPCR primer–probe sets | Nature Microbiology
Deconstructing the Polymerase Chain Reaction: Understanding and Correcting Bias Associated with Primer Degeneracies and Primer-Template Mismatches | PLOS ONE

Unreacted Labeled PCR Primers Inhibit the Signal in a Nucleic Acid Lateral Flow Assay as Elucidated by a Transport Reaction Model | ACS Measurement Science Au